History of DNA
The search for our ancestors is a national pastime — the second most popular hobby after gardening. 1 After all, you're likely here because you want to learn more about your Sinclair genealogy.
We are all related—descended from a common African ancestor who lived only 60,000 years ago. 2 For some of us, this is difficult to believe. Certainly there were those alive 2.3 million years ago when the first homo sapien walked in Oduvai Gorge in Africa. Evidence suggests, however, that many of the descendants of these early humans died off or didn’t have children.
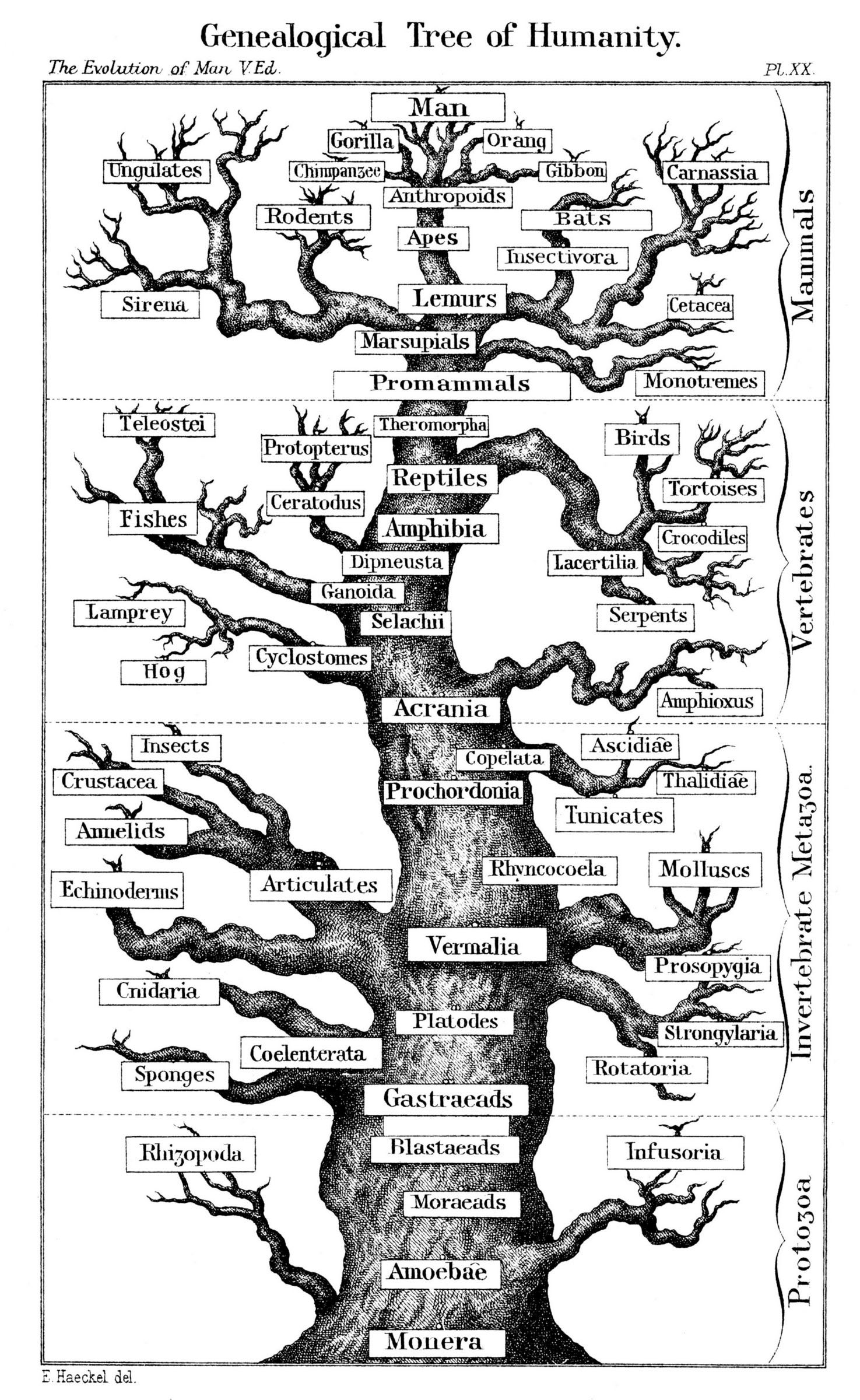
The tree of human evolution according to German evolutionary biologist Ernst Haeckel, 1891.
Today,
Speaking more specifically, we’re looking for a specific branch near the top of the genetic tree, way out on only one Haplogroup branch. Looking within a given family and trying to understand the paths of specific branches of a family (Lineages) is much more difficult and requires more than the traditional DNA test to dissect these subtle nuances. This has been our quest, to find new ways to go beyond the DNA tests currently offered, as those tests are for people who want merely to solve a question of genealogy in the 1700s.
A Brief History of Genetics
Recent History of a Recent Science
The current testing methods for identifying humans and telling us apart have evolved from techniques which had quite a different purpose. Until the 1980s, the scientific community was concerned with matching blood and tissue donors with recipients, thus reducing the rejection rate for transplant patients. Such testing was, understandably, very expensive. The advent of PCR technology made mass production commercially viable and tests were created for the hobby genealogist. PCR is a technique through which samples of DNA fragments are replicated until billions of copies are made for accurate comparison. PCR is an abbreviation for "polymerase chain reaction." (POLL'-IM-ER-ACE). This term applies to a wide variety of different DNA tests that differ in reliability and effectiveness. Reliabilities of each kind of PCR test need independent verification. PCR itself doesn't accomplish DNA typing, it only increases the amount of DNA available for typing. 105 Because of the power of PCR, very small samples of DNA from any part of the body can be used in a DNA test.
1930s - Serological Testing
1970s - HLA Testing
1980s - DNA Testing Using RFLP Technique
1990s - DNA Testing Using PCR Technology 104
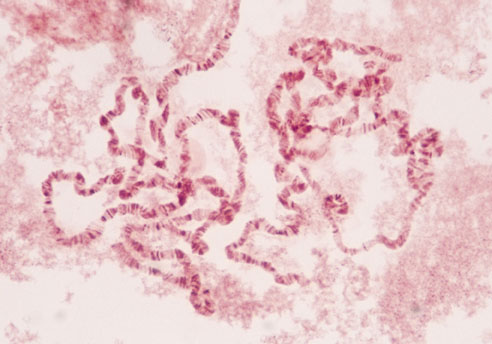
A chromosome is a very long DNA molecule, with its associated proteins, which carries
portions of the hereditary information of an organism. The human haploid genome contains
3,000,000,000 DNA nucleotide pairs.
Marketing Is Important
I run a marketing company and I know the kind of attention one must generate to achieve success. Because most of the folks exploring DNA are not scientists or academics, some do not follow good scientific method. They also don’t have a tenured position at a university at stake and so leap to conclusions.
This urgency to leap to conclusions is, to me, one of the problems inherent in the hunt for these new SNPs. For instance, Hugh Montgomery, an author on European history (who, by his own admission ‘knows nothing about DNA’) has decided that CCR5-Delta-32 is what made certain of the population of Europe into ‘God Kings,’ a title he uses to describe the descendants of his ‘Davidic Lineage.’ 11
The reading I’ve done on CCR5-delta-32 makes it very clear that there is still debate as to how old the mutation is, and that determines how much value it is for our study. If, as one researcher has suggested, it’s 127,000 years old, then it’s of no value to us in a genealogical timeframe. But because so many have books to bring to market, the tendency is to state theories as fact before the science is old enough to be reliable.
Competition Is Good
At this writing; FTDNA has just launched a new way to dissect the R1b Haplogroup. This is not surprising as so many of their results must be in this Haplogroup. We’re talking big money here. For some of the participants in the Sinclair DNA project, this new dissection will prove useful but many of us are still falling within an undeterminable “DNA soup” of the Western European Germanic tribes.
EthnoAncestry has S21. Others will surely follow. The St. Clair DNA study is closely following all the different opportunities for our project out there. We’re not wed to FTDNA. They are the largest by far, but when we see another way to understand our family connections, we’ll definitely follow it.
As more time goes by, I believe the interest in genealogy will only grow. This will drive more money into the field and companies will be even more competitive with one another to offer better answers. The R1b conundrum will continue to be a problem that these companies will want to solve because it also matches up with the largest group of immigrants in the colonies, and these folks are the ones most interested in doing their genealogy to, in my opinion, find their way home again.
Home | Contact
